A remarkably small bacterium containing fewer than 500 genes serves as the basis for one of the most detailed digital life reconstructions ever created.
Using computer technology, scientists have reconstructed in detail the complete lifecycle of this minimal living cell. They monitored that entire lifecycle from the DNA replication phase through the protein production phase and finally through the cell division phase. The simulated minimal cell behaves like a natural organism to such an extent that the predicted lifecycle of the virtual minimal cell and the actual biological minimal cell are within two minutes of each other.
The research team, led by Dr. Zan Luthey-Schulten, a chemistry professor at the University of Illinois Urbana-Champaign, documented their findings in an article published in the journal Cell.
The aim of this large project was to develop a digital model of the cellular processes in this bacterium called JCVI-syn3A, or simply Syn3A. This organism was developed at the J. Craig Venter Institute and has only those genes necessary for the essential functions of a living microorganism.

The genome of JCVI-syn3A consists of only 493 genes arranged in a single circular DNA molecule. This makes it simpler to model than any other known organism while still having the same characteristics of a living organism.
“This is a three-dimensional, fully dynamic kinetic model of a minimal life form that simulates cellular activity and development,” Dr. Luthey-Schulten told The Brighter Side of News.
The JCVI-syn3A simulation provides a comprehensive view of how a multitude of molecular interactions occur. It also shows how each molecular interaction affects the entire cellular life cycle.
The process of cell division includes gene expression, metabolism, replication of chromosomes, and reshaping of the physical structure of the cell in preparation for division. In contrast to traditional models that assume cells are homogeneous mixtures of chemicals, this model considers the fact that a real cell has a crowded, heterogeneous interior.
The researchers emphasize that cells are not simply chemical reactors. They are composed of different types of molecules that must diffuse through space and meet together for a reaction to take place.
Observing a single cell unfolding over time and space, essentially a four-dimensional view of cellular activity, has been achieved using this model. The model captures the biological processes of a cell in three spatial dimensions and time. This is equivalent to providing a four-dimensional representation of the cell’s life.
The DNA in the virtual cell behaves like a flexible polymer chain that curls, replicates, and eventually separates into two sets of daughter chromosomes. During the cell’s growth, ribosomes form and float freely within the cytoplasm. Membrane proteins also aggregate as the cell grows larger.
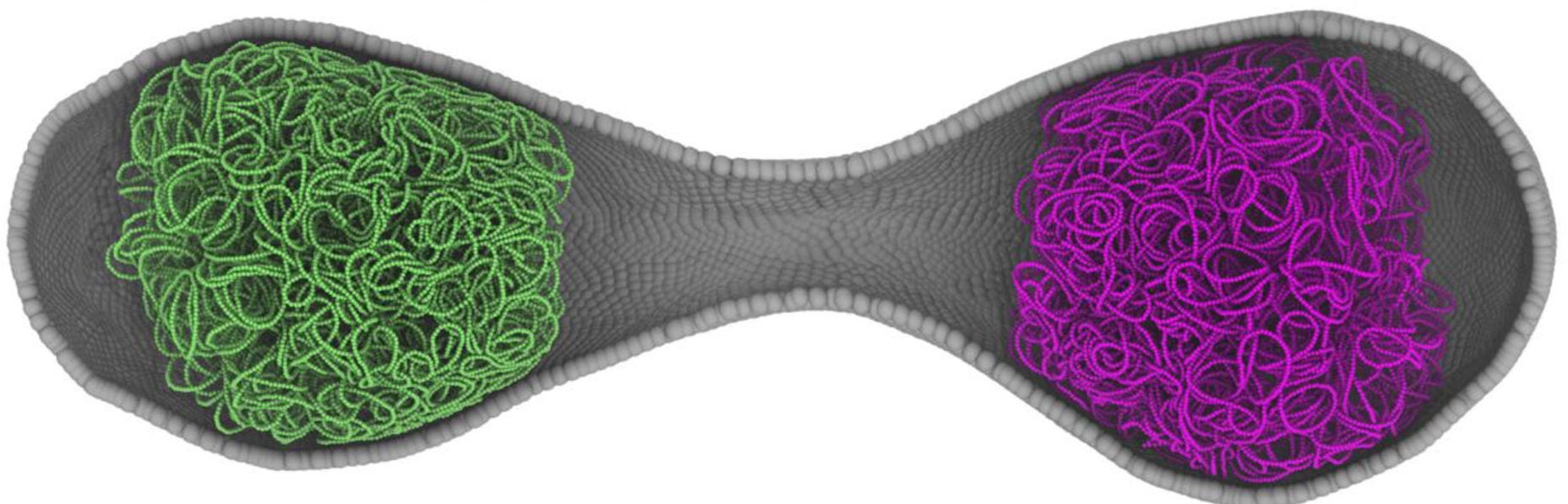
The cell’s physical shape changes during the division cycle. It initially appears spherical and then elongates prior to dividing into two daughter cells.
The high level of detail required for simulation was accomplished by combining several computational techniques. For example, the stochastic reaction methodology was used to model some metabolic processes, while ordinary differential equations were used to model others. The motions of DNA were simulated using Brownian dynamics.
Andrew Maytin, a graduate student who participated in this study, found that replication of DNA greatly reduced the speed of the simulation. He resolved this problem by running the DNA replication component of the model on a dedicated graphics processing unit. The adjustment made it possible to create a computer simulation of the complete 105-minute cell cycle using about six days of computational resources.
In addition, the simulation produced outcomes closely aligned with the actual biology of the organism.
The cell in the simulation doubled its membrane surface area during the course of the 105-minute cycle. This result matched the growth rate of Syn3A. The estimated time required to complete DNA replication in the simulated cells was approximately 51 minutes.
Researchers used experimental data in addition to modeling data to validate the findings. Imaging experiments conducted by Angad Mehta and Taekjip Ha provided confirmation that Syn3A divides symmetrically.
The experiments provided valuable insights into the configuration of the simulation. They validated the predicted cell shapes and timings.
“It is extremely difficult to create a 3D computer simulation of a single cell that has so many moving parts,” said Thornburg, a postdoctoral researcher working with the research team. “It was an incredible achievement to be able to simulate an entire cell in 3D.”
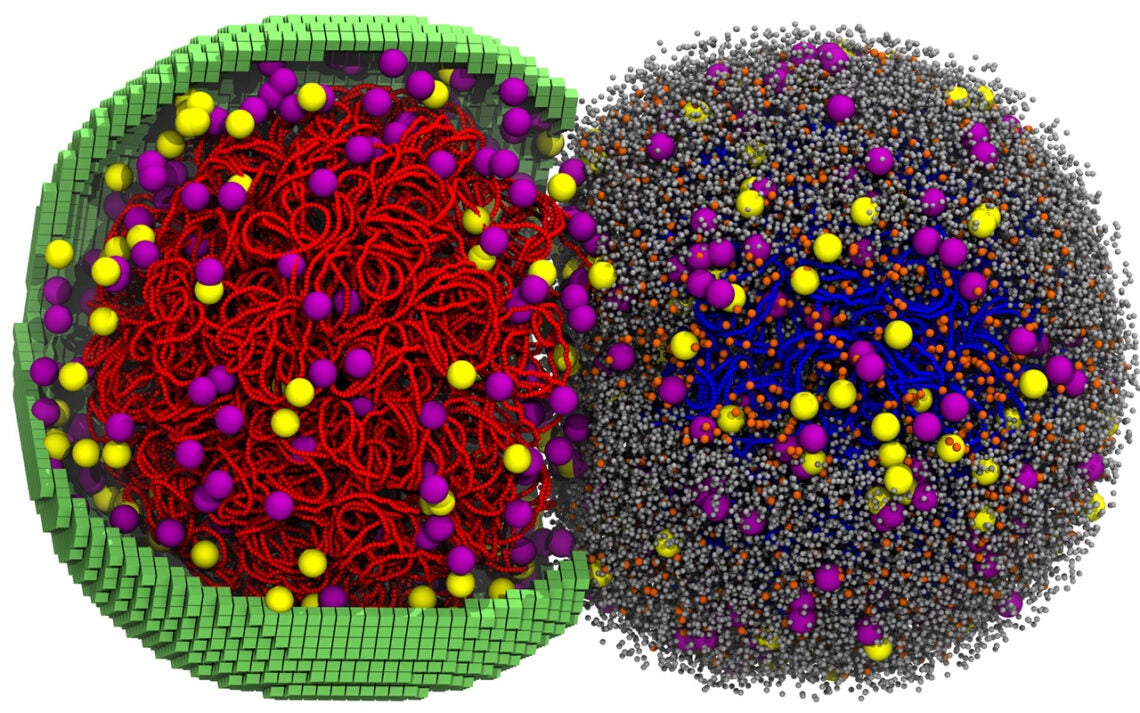
Because the interior of the simulated cell contains a large number of very small molecules, it was not feasible to visualize the entire interior simultaneously.
When researchers created animated representations of the simulation, some components had to be omitted for clarity. Eliminating cellular proteins in certain visualizations allows researchers to observe how chromosomes move through the crowded interior of the cell.
Another major finding from this model was how genetic activity, specifically transcription, is linked to metabolism. The model revealed evidence of coordinated interactions between the two.
Before RNA is synthesized, the concentration of nucleoside triphosphates supplies the necessary building blocks for transcription. Transcription rates slow as the pool of nucleoside triphosphates becomes depleted, resulting in an on-and-off pattern of transcription.
Protein synthesis is heavily dependent on ribosomal activity and the lifetime of mRNA molecules. Ribosomal activity in the simulation occurred approximately 55% of the time. The approximate percentage of RNA polymerases engaged in transcription was 70%.
As with most sophisticated models, the digital cell has limitations.
The model cannot track the movements of individual atoms. As a result, the behavior of molecules must be averaged to keep calculations manageable.
Additionally, other biological aspects have not yet been modeled. These include simultaneous translation of multiple mRNA molecules by ribosomes in polysome aggregates.
The high computational cost of this model represents another limitation. Simulating one cell cycle took between four and six days using two high-performance GPUs.
In total, the 50 simulated cells produced more than 15,000 GPU hours of computation time.
Despite these restrictions, the digital model provides an unprecedented opportunity to view biological processes related to genetics and metabolism at the cellular level.
Overall, creating a complete digital simulation of a living biological cell will give scientists new ways to study biology. Instead of running experiments on individual pathways, scientists will be able to observe how multiple processes interact simultaneously within a virtual cell.
Virtual cell simulations may ultimately predict how genetic changes affect cellular characteristics. They may also facilitate the design of synthetic organisms and provide insight into the basic principles governing all life.
The Syn3A model provides an entry point for creating more complex virtual simulations of biological organisms. This work moves researchers closer to the ultimate goal of building fully predictive biological virtual cells.
Research findings are available online in the journal Cell.
The original story “Scientists create the most detailed digital simulation of life ever attempted” is published in The Brighter Side of News.
Like these kind of feel good stories? Get The Brighter Side of News’ newsletter.
The post Scientists create the most detailed digital simulation of life ever attempted appeared first on The Brighter Side of News.