Cancer treatment has long been haunted by the same problem: how do you strike dangerous cells without hitting healthy ones nearby?
A team at the University of Geneva has built a drug-delivery system meant to answer that question with unusual precision. Instead of relying on bulky antibodies alone, the researchers used synthetic DNA strands, small binding proteins called affibodies, and aptamers to create a kind of molecular checkpoint. The treatment activates only when the right combination of markers appears on a cell’s surface.
That matters because many cancer drugs still damage healthy tissue along with tumors. Antibody-drug conjugates, or ADCs, have helped narrow that gap by linking a cancer-seeking antibody to a toxic payload. However, those therapies come with tradeoffs. Antibodies are large, which can make it harder for them to move deep into solid tumors. Most can carry only a limited number of drug molecules.
The Geneva group took a different route.
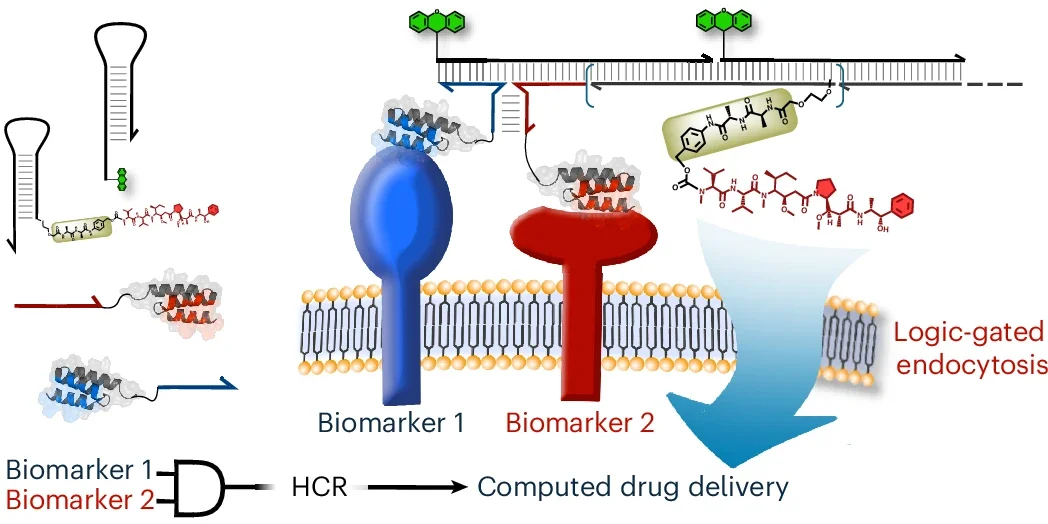
Their system uses DNA hairpins, engineered strands that remain inactive until a trigger sets off a hybridization chain reaction, or HCR. In this study, the trigger was split in two. Each half was attached to a separate cancer-targeting binder. Only when both binders latched onto matching biomarkers close together on the same cell could the DNA pieces form the full trigger. As a result, the DNA pieces could start assembling.
It worked like an AND gate in computing. Both conditions had to be met, or nothing happened.
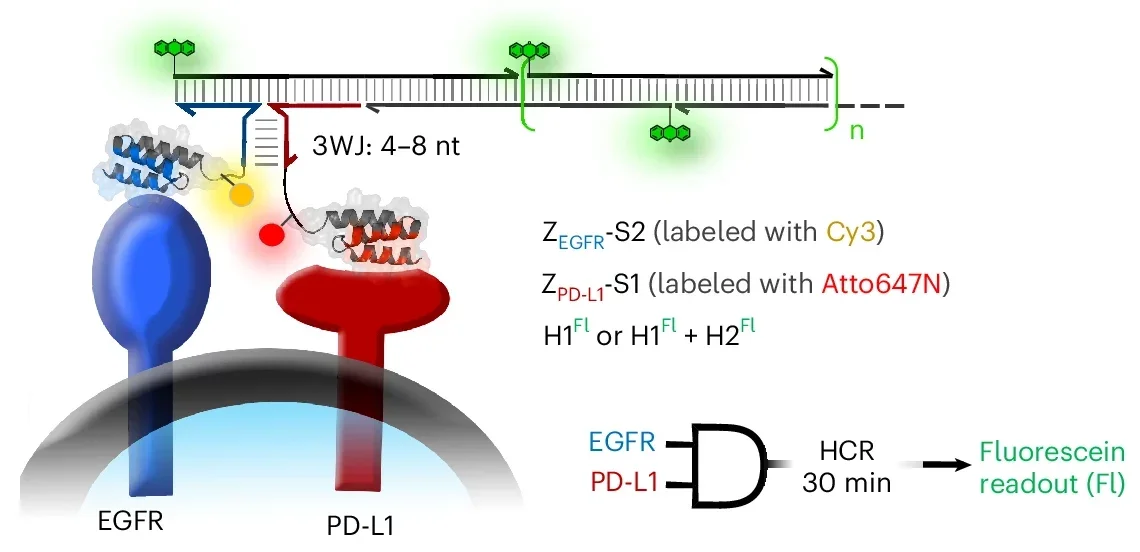
The researchers tested the approach using several biomarker pairs, including EGFR and PD-L1, and EGFR and PTK7. They also tested whether the system could respond to a single biomarker when enough copies were packed closely together on the cell surface.
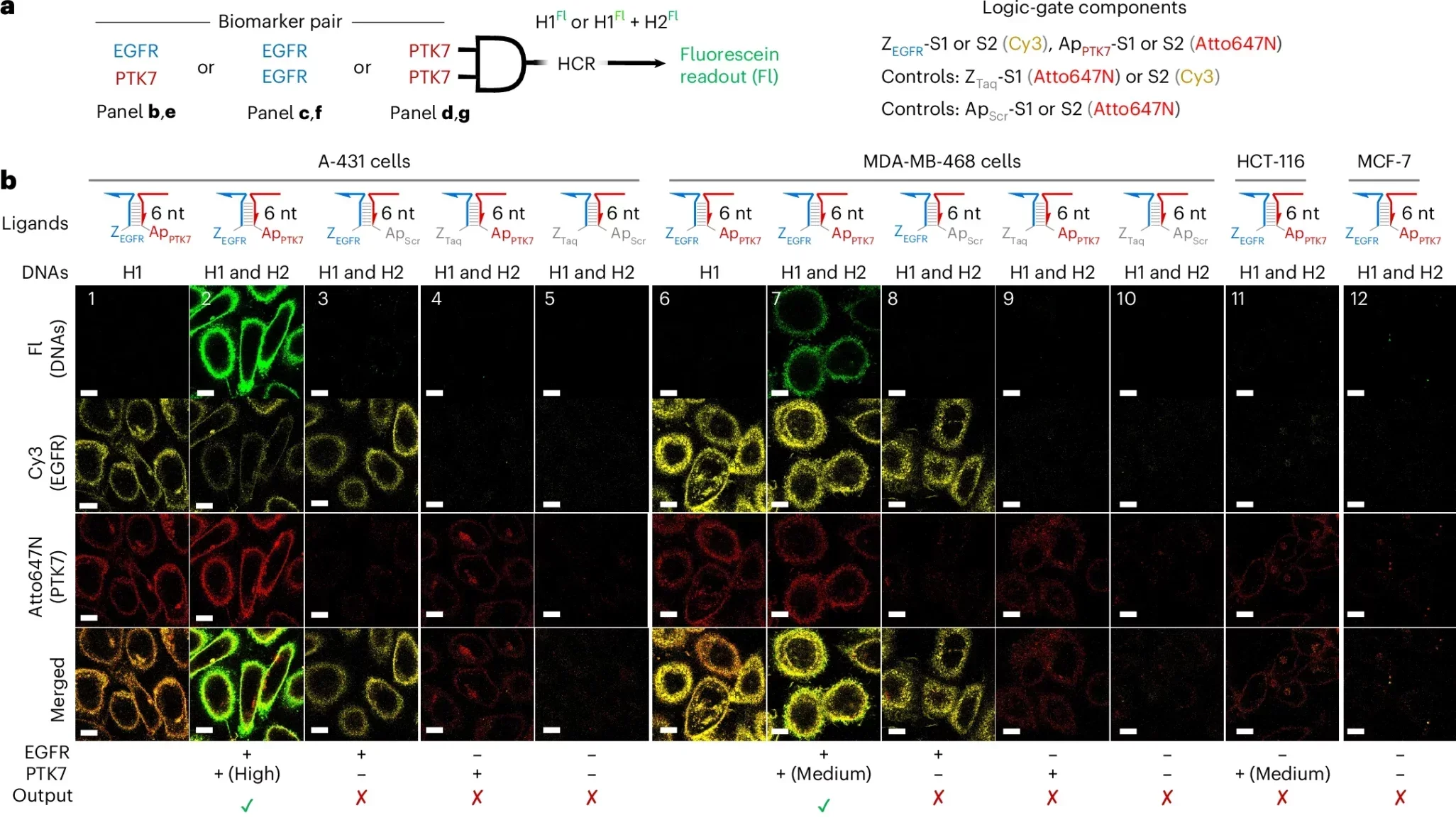
In cell experiments, the DNA reaction produced strong signal amplification only in cells carrying the right marker combinations. A-431 cells, which expressed both EGFR and PD-L1, showed a strong response. Other cell lines with only one marker, or neither, showed weak or no reaction. In another set of tests, cells carrying both EGFR and PTK7 also stood out, while control cells did not.
One result was especially striking: when the system targeted EGFR alone on A-431 cells, the median amplification reached 217 propagation steps in the chain reaction.
That amplification is central to the design. Instead of delivering a few drug molecules the way many ADCs do, the DNA system can assemble larger structures right on the cell surface. This concentrates more payload where it is needed.
“This could mark an important step forward in the evolution of medicine, with the introduction of a self-operating drug system,” said Nicolas Winssinger, a full professor in the Department of Organic Chemistry at the UNIGE School of Chemistry and Biochemistry and the study’s last author. “Until now, computers and AI have helped us design new drugs. What’s new here is that the drug itself can, in a simple way, ‘compute’ and respond intelligently to biological signals.”
To see whether the system could do more than light up cells, the team attached the chemotherapy agent monomethyl auristatin E, or MMAE, to the DNA hairpins through a cleavable linker already used in ADCs. In A-431 cells, the most heavily loaded version caused dramatic cell death. It left less than 8% viability after 48 hours. Control treatments that lacked proper cell binding did not show the same toxicity.
The system also appeared selective. In mixed-cell experiments, A-431 cells were grown alongside HeLa-GFP cells. After treatment, the A-431 population dropped sharply, while the HeLa cells kept proliferating at a rate similar to controls. The authors said that pointed to minimal bystander effects from neighboring cells that had taken up the drug.
The researchers also showed that the platform could carry more than one therapy at once. They tested a combination of MMAE and the exatecan derivative Dxd, and reported efficient delivery when the correct biomarker pair was present. That could matter because combining payloads is one way scientists try to limit drug resistance.
Then there was another twist. The same DNA assembly was able to recruit antibodies to targeted cells. In those experiments, the HCR product captured an anti-fluorescein monoclonal antibody at the cell surface. This suggests the platform might eventually be used not just for drug delivery, but also for immune-based strategies.
Not everything is solved. The study notes that the DNA used here has limited metabolic stability in plasma, with a half-life of about two hours. An unmodified aptamer used to target PTK7 also remains a limitation. The researchers point to possible fixes, including mirror-image DNA and chemical modifications that improve aptamer stability.
Even so, the study lays out a flexible framework: small synthetic parts, modular biomarker selection, and drug loading that can be tuned more freely than many protein-based systems.
This work suggests future cancer drugs could be programmed to act only when a specific mix of tumor signals appears, rather than simply recognizing one target and releasing a toxin.
That could help reduce collateral damage to healthy tissue, improve delivery inside solid tumors, and make it easier to combine multiple therapies in one treatment.
Beyond cancer, the same logic-based DNA system may also support more adaptable medicines that respond to biological cues inside the body.
Research findings are available online in the journal Nature Biotechnology.
The original story “New smart drugs precisely target and kill cancer cells” is published in The Brighter Side of News.
Like these kind of feel good stories? Get The Brighter Side of News’ newsletter.
The post New smart drugs precisely target and kill cancer cells appeared first on The Brighter Side of News.