Humans have about 3 billion DNA bases in their genetic makeup. However, most of it does not encode for protein. In the last few decades, scientists have come to realize that while much of the non-coding portion of the genome was once thought to be irrelevant, there is now overwhelming evidence that this segment plays an important role in regulating when and how genes switch on and off.
A new study conducted by Bar-Ilan University supports this finding. It reveals an example of how impactful a single change to this regulatory region can be, and how significantly one nucleotide alteration can affect an organism’s phenotype.
In an article published in Nature Communications, researchers found that inserting one nucleotide into the regulatory region of a mouse’s genome caused XX mice that normally develop as females to develop as males with testes and male genitalia. This change occurred outside the protein-coding region. It was located roughly 500,000 bases away from the gene it regulates.
“This is an extraordinary finding, particularly because just a minute alteration — a single DNA letter from approximately 2.8 billion total nucleotides — can lead to such a dramatic developmental change,” states Dr. Nitzan Gonen of Bar-Ilan University’s Goodman Faculty of Life Sciences. “It demonstrates how non-coding DNA has the capability to influence mammalian development and affect diseases.”
To better understand this effect, it is necessary to look at how mammals determine sex at the molecular level. The genetic sex of a mammal is not solely determined by the chromosomes present in the genome. It is not an automatic process, as additional signals are involved.
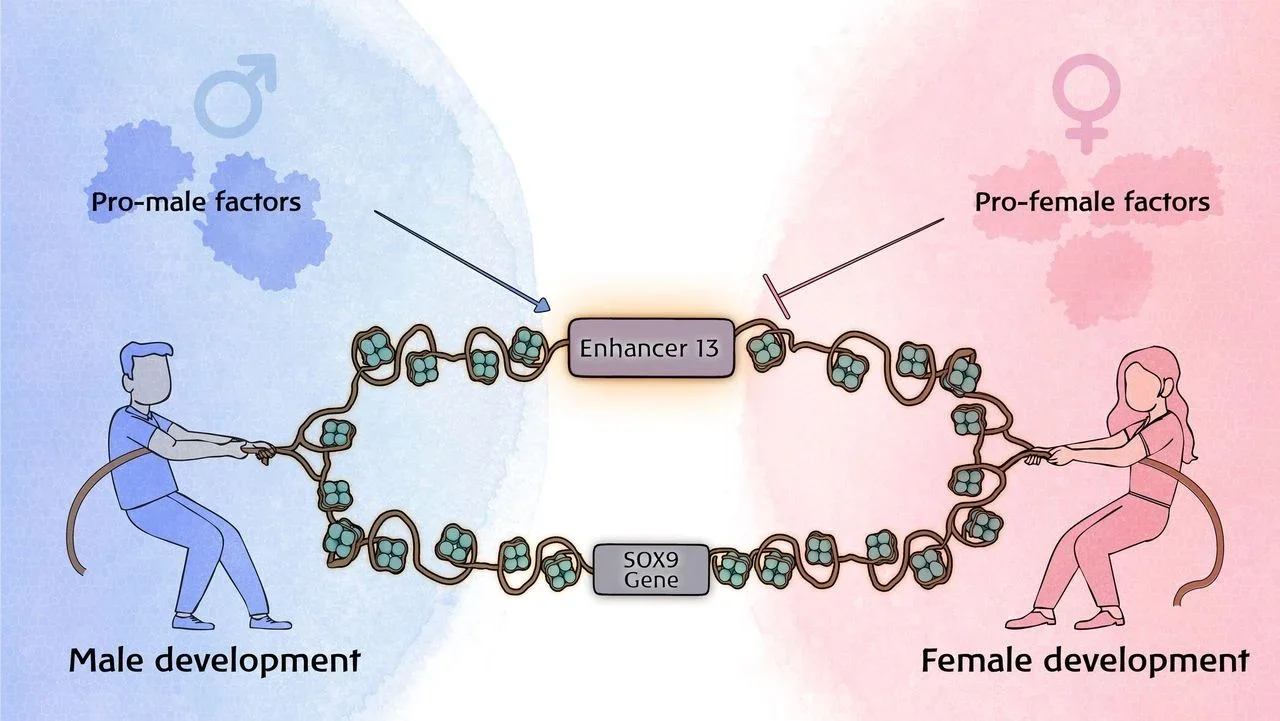
Competition between two opposing developmental programs lies at the core of this process. One program leads to the development of testes, while the other leads to ovaries. The relative strength of these competing programs is determined by molecular signals and their abundance during a critical early stage of embryonic development.
The gene Sox9 is a key player in testicular development. When Sox9 is activated, it initiates a cascade of molecular events that leads to the formation of Sertoli cells and the organization of testicular tubules. It directs the developing gonad toward the male pathway.
Conversely, for the gonad to develop into an ovary, Sox9 expression must remain suppressed. In XY individuals, Sox9 is activated by Sry gene expression located on the Y chromosome. This initiates the testicular developmental pathway. In XX individuals, multiple pro-female factors cooperate to suppress Sox9 and allow the ovarian pathway to proceed.
The Bar-Ilan research group previously demonstrated that Enh13, a 557 base-pair regulatory sequence located hundreds of kilobases from Sox9, acts as a “master switch” in XY embryos. It provides the enhancer function necessary to activate Sox9 expression.
The present study reveals that Enh13 has an additional function. It also helps suppress Sox9 expression through the action of the ovarian developmental program in XX embryos.
The researchers created two different small mutations in Enh13 using CRISPR genome editing technology. One mutation was a three-letter deletion, and the other was a one-letter insertion. Despite these differences, the XY embryos with either mutation developed as females.
Both mutations occurred at the same location within Enh13. This region is where Sox9 binds to itself to strengthen its own expression during testis development.
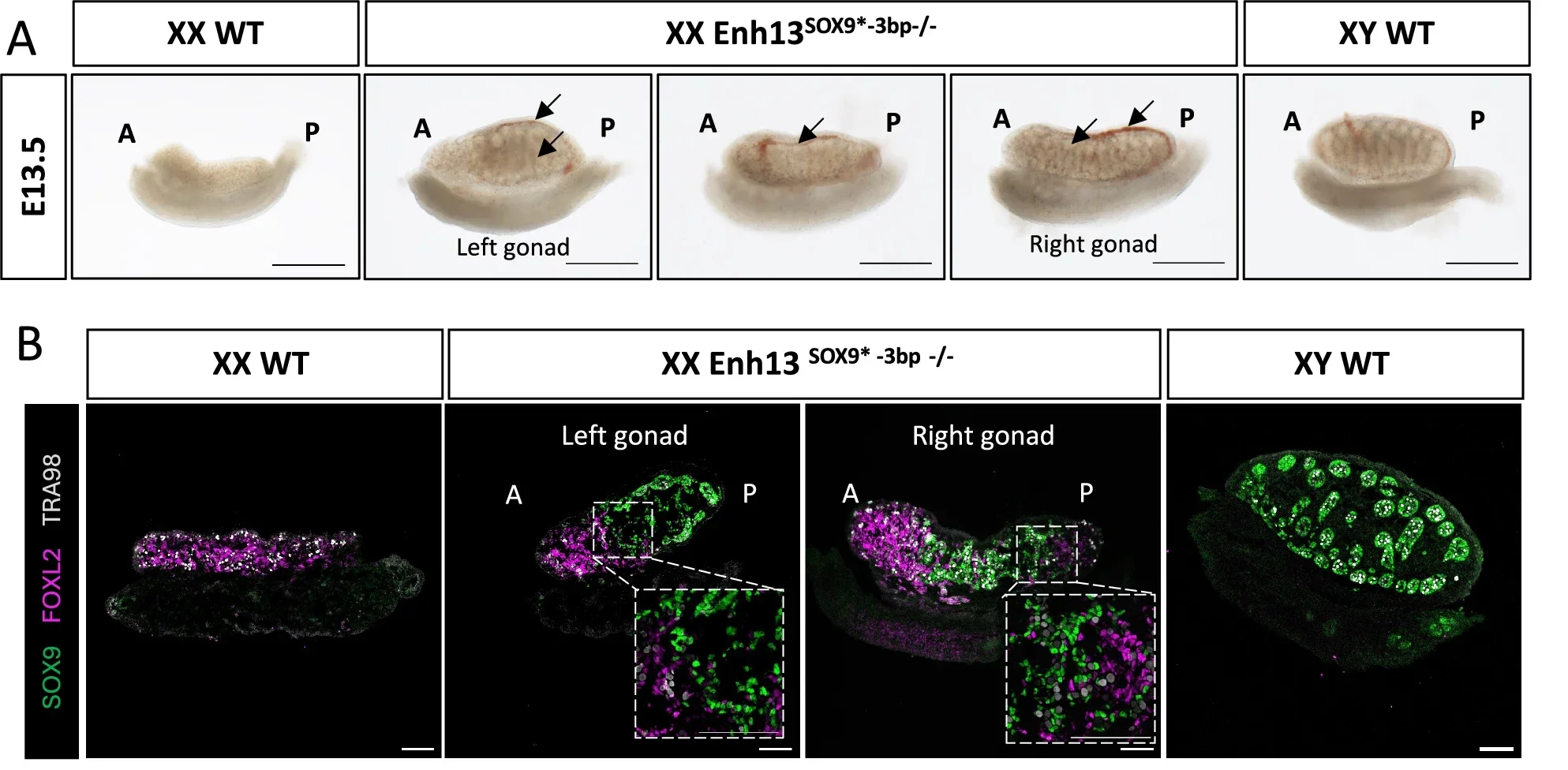
Experiments using XX mice with either mutation resulted in adult mice that appeared male. They developed testes, male internal organs, and male external genitalia. As expected, the testes did not contain sperm because sperm production requires Y chromosome genes.
The embryos of the XX mice were analyzed to understand how their development proceeded. The gonads initially formed as ovotestes, which contain both ovarian and testicular tissue. This occurred before the completion of organ development.
During development, the testicular components became dominant. The animals were ultimately born with fully developed male anatomy.
“Our study demonstrates that studying genes alone is not sufficient,” says Elisheva Abberbock, the lead author of the research. “Important pathogenic mutations may actually reside in the non-coding region of the genome, which regulates gene expression rather than encodes protein products.”
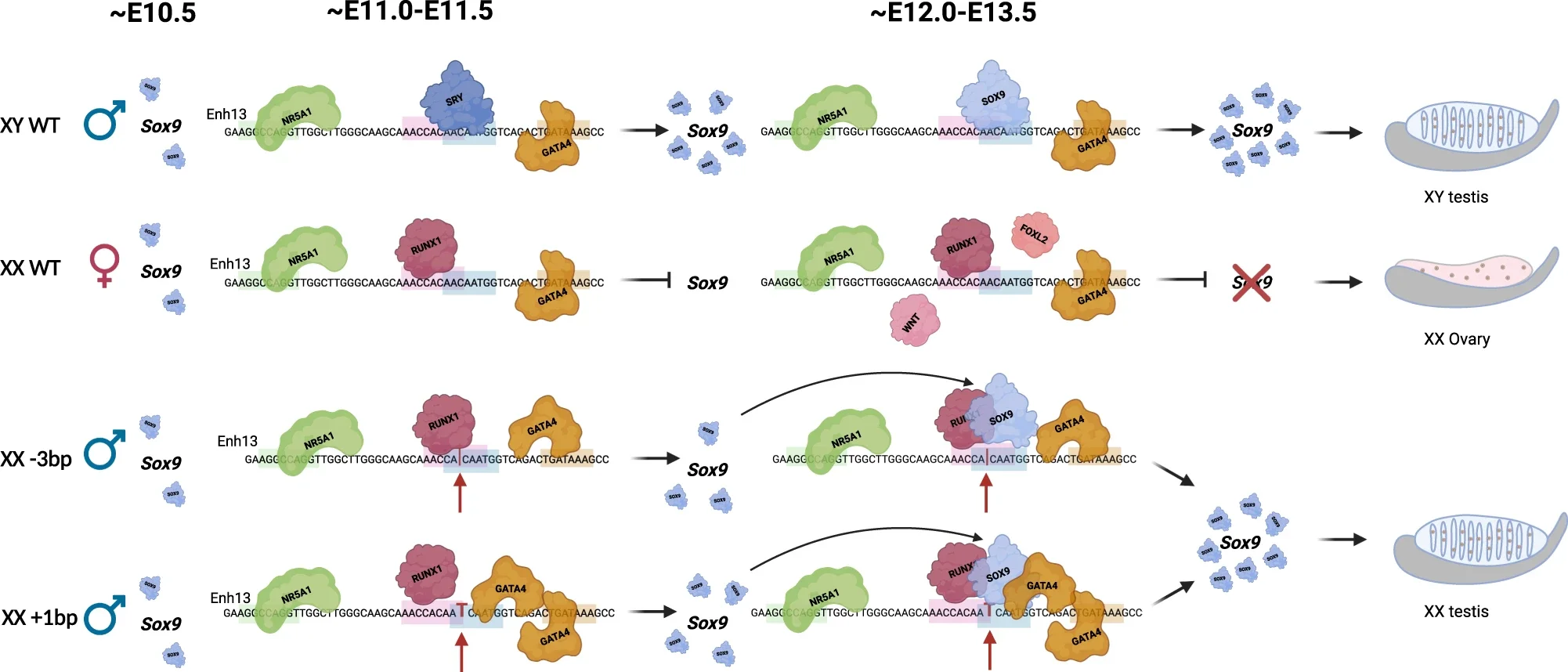
The next stage of the study focused on how such small mutations produced large effects. The mechanism did not simply involve stronger binding of Sox9 to DNA. Instead, it involved competition between Sox9 and other transcription factors for the same regulatory region, Enh13.
Among the binding sites identified were those for SOX9, RUNX1, NR5A1, and GATA4. RUNX1 is associated with promoting ovarian development. NR5A1 and GATA4 are present in embryonic gonads of both sexes. There is significant overlap between the binding locations of SOX9 and RUNX1.
The researchers found that in normal females, Sox9 is repressed by RUNX1 and other pro-female proteins. These proteins bind to nearby locations along the DNA, often at conserved distances from one another.
This spacing appears to be important. The distance between binding sites is conserved across mammals, suggesting that it plays a functional role. When mutations change this spacing, they alter how proteins interact with each other when bound to DNA.
While the mutations do not block repression entirely, they appear to shift the balance. The repressive configuration is converted into a weakly activating one. Once Sox9 activity crosses a critical threshold, an amplification loop increases its expression, initiating testis development.
The one-base-pair insertion produces an additional effect. It creates a new binding site for GATA4 adjacent to the RUNX1 site. This leads to a stronger activation signal compared to the three-base-pair deletion.
As a result, the insertion produces a more pronounced male phenotype. It also accelerates the process of testis differentiation.
Approximately one in 4,000 births involves Disorders of Sex Development (DSD). These conditions include cases where chromosomal sex, gonadal development, or genital development does not follow typical patterns.
More than 50 percent of children born with DSD do not receive a genetic diagnosis through sequencing of protein-coding genes. The findings from the Bar-Ilan study suggest that mutations in non-coding regions may explain some of these cases.
The researchers found that individuals with 46,XX DSD, who develop male characteristics, often carry small duplications in the genomic region containing Enh13. The mechanism behind these duplications has not been fully studied.
However, the findings suggest that these duplications may alter the regulatory environment. This could shift the balance of Sox9 activation in XX individuals.
The study also raises the possibility that similar regulatory mechanisms exist in other species. For example, DSD has been observed in pigs where XX individuals lacking the Sry gene still develop male traits. These cases have been mapped to the Sox9 region, although not directly to the gene itself or its known enhancers.
The findings reflect a broader principle in regulatory genomics. The arrangement of transcription factor binding sites within an enhancer is highly specific. It is not random.
Even small changes in enhancer sequences can alter the three-dimensional arrangement of proteins. This can change how genes are regulated in significant ways.
From a clinical perspective, the research highlights a limitation of current genetic testing. Sequencing only the coding regions of the genome may miss important mutations.
The non-coding portion of the genome is much larger and remains poorly understood. The Bar-Ilan research team plans to search for additional regulatory regions involved in sex determination and other developmental disorders.
The Enh13 system provides a useful model for studying these effects. Mutations in this region do not cause lethality. They produce clear and observable outcomes, making it easier to study how small sequence changes affect gene regulation.
Research findings are available online in the journal Nature Communications.
The original story “Flipping a single DNA letter can trigger complete sex reversal” is published in The Brighter Side of News.
Like these kind of feel good stories? Get The Brighter Side of News’ newsletter.
The post Flipping a single DNA letter can trigger complete sex reversal appeared first on The Brighter Side of News.
Leave a comment
You must be logged in to post a comment.